Can we use pipelines developed for human NGS analysis and quickly apply them for viral analysis? With ebolavirus being in the news, it seemed like a good time to try. Just as with a human sequencing project, it’s helpful if we have a good reference genome. The NCBI has four different ebola strain reference files located at their ftp:
Can we use pipelines developed for human NGS analysis and quickly apply them for viral analysis? With ebolavirus being in the news, it seemed like a good time to try. Just as with a human sequencing project, it’s helpful if we have a good reference genome. The NCBI has four different ebola strain reference files located at their ftp:
Remote directory: /genomes/Viruses/*
Accession: NC_002549.1 : 18,959 bp linear cRNA
Accession: NC_014372.1 : 18,935 bp linear cRNA
Accession: NC_004161.1 : 18,891 bp linear cRNA
Accession: NC_014373.1 : 18,940 bp linear cRNA
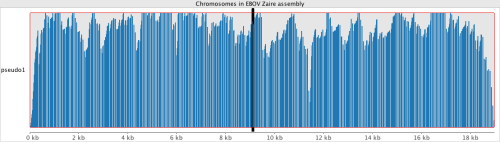
Currently everything that’s happened in West Africa looks to match best with NC_002549.1, the Zaire strain. The Broad Institute began metagenomic sequencing from human serum this summer and the data can be accessed here (Accession: PRJNA257197). We can take some of these datasets and map them to NC_002549.1. The datasets are in .sra format, and must be extracted using fastq-dump.
We can see that the data maps really well to this strain. All four of the reference genomes above were indexed with a new build of bwa(0.7.10-r876-dirty git clone https://github.com/lh3/bwa.git). Because EBOV genomes are so small, compared to humans, the only alignment algorithm which seemed suitable within bwa, was mem.
EBOV mokas$ ./bwa/bwa mem Zaire_ebolavirus_uid14703.fa SRR1553514.fastq > SRR1553514.sam
[M::main_mem] read 99010 sequences (10000010 bp)...
[M::mem_process_seqs] Processed 99010 reads in 8.988 CPU sec, 9.478 real sec
[M::main_mem] read 99010 sequences (10000010 bp)...
[M::mem_process_seqs] Processed 99010 reads in 8.964 CPU sec, 9.671 real sec
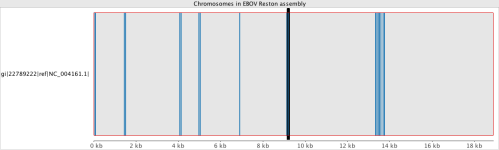
If we take the same SRA data and try to map it to some of the other strain references, e.g. the Reston Virginia strain from 1989, it can help give a rough idea of how closely related the 2014 incident is.
It can be seen that apart from a few highly conserved regions where many reads align, the coverage map indicates that the data collected in West Africa and sequenced on the Illumina HiSeq2500 does not match to NC_004161.1. There were still approximately 500 variants with the Zaire reference on the 2014 samples, showing a good amount differences, considering the entire genome is only 18,000bp.
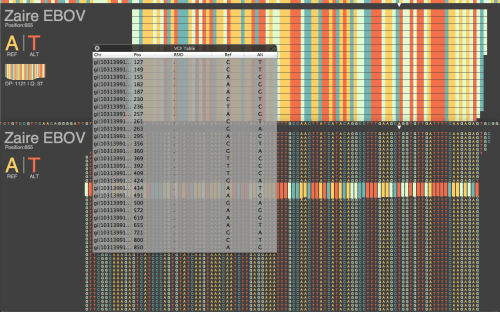
LucidAlign genome browser comparing the two strains
All of this is, of course, good news. We can take sequencing data of new EBOV strains and apply slightly modified pipelines to get meaningful results. And with the Ion PGM now being FDA approved means data can be generated in nearly 3 hours, with Federal approval. There have even been some publications which show that the protein VP24 can stop EBOV all together [DOI: 10.1086/520582] with the structures available for analysis as well. So, it looks like it’s all coming up humanity, our capabilities are there, and with proper resources this scary little bug can be a thing of history.
There have even been some publications which show that the protein VP24 can stop EBOV all together [DOI: 10.1086/520582] with the structures available for analysis as well. So, it looks like it’s all coming up humanity, our capabilities are there, and with proper resources this scary little bug can be a thing of history.