Discovering genomic variations within a single individual, which is also the underlying factor in a previously undiagnosed pathology, can be thought of as a anomaly detection problem. Colloquially referred to as the needle in a haystack.
The NCBI’s human reference genomes allows for the largest filter, enabling identification of initial variants. Next, alternate loci patches to the primary build of the human reference genome, accounting for large regions of variability, will reduce the number of variants, which will still remain too large for efficient annotation. An additional resource taps into SNP databases. The NCBI’s dbSNP provides a large set of SNP locations, meanwhile The National Cancer Institute also contains a large curated database of SNPs which are placed within three categories: Confirmed, Validated, and Candidate SNPs.
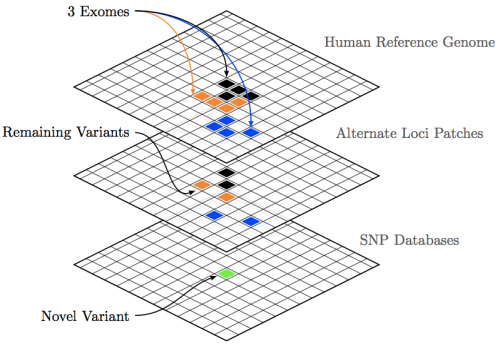
Shown in the figure above are three exomes which, after comparison with the primary human reference build contain large variant sets. These are then passed on to alternate loci, and finally SNP filters. The end result being discovery of novel variants, which may be responsible for idiopathic indications.